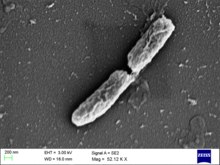
Shigella flexneri
| Shigella flexneri | |
|---|---|
| |
| Shigella flexneri | |
| Scientific classification | |
| Domain: | Bacteria |
| Phylum: | Pseudomonadota |
| Class: | Gammaproteobacteria |
| Order: | Enterobacterales |
| Family: | Enterobacteriaceae |
| Genus: | Shigella |
| Species: | S. flexneri
|
| Binomial name | |
| Shigella flexneri Castellani & Chalmers 1919
| |
Shigella flexneri is a species of Gram-negative bacteria in the genus Shigella that can cause diarrhea in humans. Several different serogroups of Shigella are described; S. flexneri belongs to group B. S. flexneri infections can usually be treated with antibiotics, although some strains have become resistant. Less severe cases are not usually treated because they become more resistant in the future.[1] Shigella are closely related to Escherichia coli, but can be differentiated from E.coli based on pathogenicity, physiology (failure to ferment lactose or decarboxylate lysine) and serology.[2]
Discovery
The species was named after the American physician Simon Flexner; the genus Shigella is named after Japanese physician Kiyoshi Shiga, who researched the cause of dysentery. Shiga entered the Tokyo Imperial University School of Medicine in 1892, during which he attended a lecture by Dr. Shibasaburo Kitasato. Shiga was impressed by Dr. Kitasato's intellect and confidence, so after graduating, he went to work for him as a research assistant at Institute for Infectious Diseases. In 1897, Shiga focused his efforts on what the Japanese referred to as a "Sekiri" (dysentery) outbreak. These epidemics were detrimental to the Japanese people and occurred often in the late 19th century. The 1897 sekiri epidemic affected >91,000, with a mortality rate of >20%.[3] Shiga studied 32 dysentery patients and used Koch's Postulates to successfully isolate and identify the bacterium causing the disease. He continued to study and characterize the bacterium, identifying its methods of toxin production i.e. Shiga Toxin, and worked tirelessly to create a vaccine for the disease.
Characterization
Morphology
Shigella flexneri is a rod shaped, nonflagellar bacterium that relies on actin-based motility. It produces the protein actin in a swift and continuous fashion to propel itself forward within and between the host's cells.[4] This bacterium is gram-negative, non-spore forming Shigella from serogroup B. There are 6 serotypes within this serogroup.[2]
Serotype
Shigella flexneri belongs to group B (i.e. agglutinate with B antisera) which further subclassified by six type-specific and four group-specific antisera. Until now at least 23 different subserotypes have been identified and reported.[5] PCR based molecular serotyping technique are now available targeting wzx1-5 (All except serotype 6) and gtr genes or wzx6 (Only serotype 6).[6]
| Serotype | Previously designated name | Type antigen specific antisera (MASF) | MASF | Group antigen specific antisera (MASF) | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| I | II | IV-2 | V | VI | Ic | B | Y-5 | 6 | 7,8 | IV-I | ||
| S. flexneri 1a | 1a | + | + | + | ||||||||
| S. flexneri 1b | 1b | + | + | + | ||||||||
| S. flexneri 1d | 1d | + | + | + | ||||||||
| S. flexneri 2a | 2a | + | + | + | ||||||||
| S. flexneri 2b | 2b | + | + | + | ||||||||
| S. flexneri 3a | 3a | + | + | + | ||||||||
| S. flexneri 3b | 3b | + | + | |||||||||
| S. flexneri 4a | 4a | + | + | + | ||||||||
| S. flexneri 4b | 4b | + | + | + | ||||||||
| S. flexneri 4c | 4c | + | + | + | ||||||||
| S. flexneri 4d | type4 | + | + | + | ||||||||
| S. flexneri 4e | 4a, 4av | + | + | + | + | |||||||
| S. flexneri 5a | 5a | + | + | + | ||||||||
| S. flexneri 5b | 5b | + | + | + | ||||||||
| S. flexneri 6a | type 6 | + | + | |||||||||
| S. flexneri 6b | type 6 | + | + | + | ||||||||
| S. flexneri 7a | 1c | + | + | |||||||||
| S. flexneri 7b | 1c+6 | + | + | + | ||||||||
| S. flexneri Xa | X | + | + | |||||||||
| S. flexneri Xb | Xv | + | + | + | ||||||||
| S. flexneri Ya | Y | + | + | |||||||||
| S. flexneri Yb | Yv | + | + | + | ||||||||
| S. flexneri Z | 4X, 4s | + | + | |||||||||
Invasion
Shigella flexneri is an intracellular bacterium that infects the epithelial lining of the mammalian intestinal tract. This bacterium is acid tolerant and can survive conditions of pH 2. Thus, it is able to enter the mouth of its host and survive passage through the stomach to the colon.[7] Once inside of the colon, S. flexneri can penetrate the epithelium in three ways: 1) The bacterium can alter the tight junctions between the epithelial cells, allowing it to cross into the sub-mucosa. 2) It can penetrate the highly endocytic M cells that are dispersed in the epithelial layer and cross into the sub-mucosa. 3) After reaching the sub-mucosa, the bacteria can be phagocytosed by macrophages and induce apoptosis, cell death. This releases cytokines that recruit polymorphonuclear cells (PMN) to the sub-mucosa. S. flexneri still in the lumen of the colon traverse the epithelial lining as the PMNs cross into the infected area. The influx of PMN cells across the epithelial layer in response to Shigella disrupts the integrity of the epithelium allowing lumenal bacteria to cross into the sub-mucosa in an M-cell independent mechanism.[8] S. flexneri uses these three methods to reach the sub-mucosa to penetrate the epilithelial cells from the basolateral side. The bacterium has four known invasion plasmid antigens: IpaA, IpaB, IpaC, and IpaD. When S. flexneri makes contact with the basolateral side of an epithelial cell, IpaC and IpaB are fused together to make a pore in the epithelial cell membrane. It then uses a type-III secretion system (T3SS) to insert the other Ipa proteins into the cytoplasm of the epithelial cell.[8] S. flexneri can pass to neighboring epithelial cells by using its own outer membrane protein, IcsA, to activate the host's actin assembly machinery. The IcsA protein is first localized to one pole of the bacterium where it will then bind with the host's protein, Neural Wiskott-Aldrich Syndrome Protein (N-WASP). This IcsA/N-WASP complex then activates the Actin-related protein (Arp) 2/3 Complex. Arp 2/3 Complex is the protein responsible for rapidly initiating actin polymerization and propelling the bacteria forward.[8][2][9] When S. flexneri reaches the adjoining membrane, it creates a protrusion into the neighboring cell's cytoplasm. The bacteria becomes surrounded by two layers of cellular membrane. It then uses another IpaBC complex to make a pore and enter the next cell. VacJ is a protein that is also needed by S. flexneri to exit the protrusion. Its exact function is still being studied but it is known that intercellular spread is greatly impaired without it.[8][10] Bacterial replication within the epithelial cell is detrimental to the cell but it is proposed that epithelial cell death is largely due to the host's own inflammatory response.[8]
Genetics
The genome of S. flexneri and Escherichia coli are nearly indistinguishable at the species level. S. flexneri has a circular chromosome with 4,599,354 base pairs. It is smaller than that of E. coli but the genes are similar. S. flexneri has about 4,084 known genes in the genome. The extensive similarity between E. coli and S. flexneri is proposed to be due to horizontal transfer. All of the genes needed for S. flexneri to invade the epithelial lining of the colon are found on a virulence plasmid called pINV. The genome of pINV is highly conserved between subspecies of S. flexneri. S. flexneri also has two other small multicopy plasmids, but some strains of S. flexneri have more plasmids that are suspected to confer antibiotic resistance.[11] Some strains of S. flexneri have resistance to the antibiotics streptomycin, ampicillin, or trimethoprim.[12] It has been found that chloramphenicol, nalidixic acid, and gentamicin are still effective antibiotics for some strains.[13]
Metabolism
Shigella flexneri is a heterotroph. It utilizes the Embden-Meyerhof-Parnas (EMP), Entner-Doudoroff (ED), or pentose phosphate pathway (PPP) to metabolize sugars. The products of these pathways then feed into the Citric Acid Cycle (TCA). S. flexneri can metabolize glucose and pyruvate. Supplemented pyruvate allows for the most growth and is believed to be the preferred carbon source. Pyruvate could be supplied by the cell's own metabolism or taken from the host cell. S. flexneri is a facultative anaerobe that is able to perform mixed-acid fermentation of pyruvate.[14][2] S. flexneri is unable to ferment lactose.[2] This bacterium grows optimally at 37 °C but can grow in temperatures as low as 30 °C.[13]
Small RNA
Bacterial small RNAs play important roles in many cellular processes. RnaG and RyhB sRNAs have been well studied in S. flexneri.[15] Ssr1 sRNA, which could play role in resistance to acidic stress and regulation of virulence was shown to exist only in Shigella.[16]
Ecology
Infectious cycle
Shigella flexneri contains a virulence plasmid that codes for three virulence factors: a type-3 secretion system (T3SS), invasion plasmid antigen proteins (IPA proteins), and IcsA (used for cell-to-cell spread).[17]
Upon infection, S. flexneri injects the host cell cytoplasm with ipa proteins using the T3SS—a needle-and-syringe-like apparatus common to many Gram-negative pathogens. These ipa proteins induce "membrane ruffling" by the host cell. Membrane ruffling creates membrane pockets which capture and engulf the bacteria. Once inside, S. flexneri uses host cell actin for propulsion to move directly from cell to cell using a cellular mechanism known as paracytophagy,[18][19] similarly to the bacterial pathogen Listeria monocytogenes.
Shigella flexneri is able to inhibit the acute inflammatory response in the initial stage of infection[20] by using an effector protein, OspI, which is encoded by ORF169b on the Shigella large plasmid and secreted by the type III secretion system. It dampens the inflammatory response during bacterial invasion by suppressing the TNF-α-receptor-associated factor 6 (TRAF6)-mediated signalling pathway.[20] OspI has glutamine deamidase activity, and is able to selectively deaminate glutamine at position 100 in UBC13 to glutamate, and this results in a failure of the E2 ubiquitin-conjugating activity which is required for TRAF6 activation.[20]
References
- ^ Ryan KJ; Ray CG; Sherris JC, eds. (2004). Sherris Medical Microbiology (4th ed.). New York: McGraw-Hill. ISBN 978-0-8385-8529-0. LCCN 2003054180. OCLC 52358530.
- ^ a b c d e Hale, Thomas L.; Keusch, Gerald T. (1996), Baron, Samuel (ed.), "Shigella", Medical Microbiology (4th ed.), University of Texas Medical Branch at Galveston, ISBN 978-0-9631172-1-2, PMID 21413292, retrieved 2020-04-23
- ^ Trofa, Andrew F.; Ueno-Olsen, Hannah; Oiwa, Ruiko; Yoshikawa, Masanosuke (1999-11-01). "Dr. Kiyoshi Shiga: Discoverer of the Dysentery Bacillus". Clinical Infectious Diseases. 29 (5): 1303–1306. doi:10.1086/313437. ISSN 1058-4838. PMID 10524979.
- ^ Goldberg, Marcia B. (December 2001). "Actin-Based Motility of Intracellular Microbial Pathogens". Microbiology and Molecular Biology Reviews. 65 (4): 595–626. doi:10.1128/MMBR.65.4.595-626.2001. ISSN 1092-2172. PMC 99042. PMID 11729265.
- ^ Shahnaij, Mohammad; Latif, Hasan A.; Azmi, Ishrat J.; Amin, Mohammed Badrul; Luna, Sharmin J.; Islam, Mohammad Aminul; Talukder, Kaisar Ali (2018). "Characterization of a serologically atypical Shigella flexneri Z isolated from diarrheal patients in Bangladesh and a proposed serological scheme for Shigella flexneri". PLOS ONE. 13 (8): e0202704. Bibcode:2018PLoSO..1302704S. doi:10.1371/journal.pone.0202704. ISSN 1932-6203. PMC 6108489. PMID 30142163.
- ^ Brengi, Silvina P.; Sun, Qiangzheng; Bolaños, Hilda; Duarte, Francisco; Jenkins, Claire; Pichel, Mariana; Shahnaij, Mohammad; Sowers, Evangeline G.; Strockbine, Nancy; Talukder, Kaisar A.; Derado, Gordana; Viñas, María Rosa; Kam, Kai Man; Xu, Jianguo; Onderdonk, Andrew B. (2019). "PCR-Based Method for Shigella flexneri Serotyping: International Multicenter Validation". Journal of Clinical Microbiology. 57 (4): e01592-18. doi:10.1128/JCM.01592-18. ISSN 0095-1137. PMC 6440786. PMID 30700505.
- ^ Bagamboula, C. F.; Uyttendaele, M.; Debevere, J. (2002). "Acid tolerance of Shigella sonnei and Shigella flexneri". Journal of Applied Microbiology. 93 (3): 479–486. doi:10.1046/j.1365-2672.2002.01714.x. ISSN 1365-2672. PMID 12174047. S2CID 44572279.
- ^ a b c d e Jennison, Amy V.; Verma, Naresh K. (2004-02-01). "Shigella flexneri infection: pathogenesis and vaccine development". FEMS Microbiology Reviews. 28 (1): 43–58. doi:10.1016/j.femsre.2003.07.002. ISSN 0168-6445. PMID 14975529.
- ^ Egile, Coumaran; Loisel, Thomas P.; Laurent, Valérie; Li, Rong; Pantaloni, Dominique; Sansonetti, Philippe J.; Carlier, Marie-France (1999-09-20). "Activation of the Cdc42 Effector N-Wasp by the Shigella flexneri Icsa Protein Promotes Actin Nucleation by Arp2/3 Complex and Bacterial Actin-Based Motility". Journal of Cell Biology. 146 (6): 1319–1332. doi:10.1083/jcb.146.6.1319. ISSN 0021-9525. PMC 2156126. PMID 10491394.
- ^ Carpenter, Chandra D.; Cooley, Benjamin J.; Needham, Brittany D.; Fisher, Carolyn R.; Trent, M. Stephen; Gordon, Vernita; Payne, Shelley M. (2014-02-01). "The Vps/VacJ ABC Transporter Is Required for Intercellular Spread of Shigella flexneri". Infection and Immunity. 82 (2): 660–669. doi:10.1128/IAI.01057-13. ISSN 0019-9567. PMC 3911398. PMID 24478081.
- ^ Wei, J.; Goldberg, M. B.; Burland, V.; Venkatesan, M. M.; Deng, W.; Fournier, G.; Mayhew, G. F.; Plunkett, G.; Rose, D. J.; Darling, A.; Mau, B. (2003-05-01). "Complete Genome Sequence and Comparative Genomics of Shigella flexneri Serotype 2a Strain 2457T". Infection and Immunity. 71 (5): 2775–2786. doi:10.1128/IAI.71.5.2775-2786.2003. ISSN 0019-9567. PMC 153260. PMID 12704152.
- ^ Pan, Jing-Cao; Ye, Rong; Meng, Dong-Mei; Zhang, Wei; Wang, Hao-Qiu; Liu, Ke-Zhou (2006-08-01). "Molecular characteristics of class 1 and class 2 integrons and their relationships to antibiotic resistance in clinical isolates of Shigella sonnei and Shigella flexneri". Journal of Antimicrobial Chemotherapy. 58 (2): 288–296. doi:10.1093/jac/dkl228. ISSN 0305-7453. PMID 16766536.
- ^ a b Oaks, E. V.; Wingfield, M. E.; Formal, S. B. (1985-04-01). "Plaque formation by virulent Shigella flexneri". Infection and Immunity. 48 (1): 124–129. doi:10.1128/IAI.48.1.124-129.1985. ISSN 0019-9567. PMC 261924. PMID 3884506.
- ^ Waligora, E. A.; Fisher, C. R.; Hanovice, N. J.; Rodou, A.; Wyckoff, E. E.; Payne, S. M. (2014-07-01). "Role of Intracellular Carbon Metabolism Pathways in Shigella flexneri Virulence". Infection and Immunity. 82 (7): 2746–2755. doi:10.1128/IAI.01575-13. ISSN 0019-9567. PMC 4097621. PMID 24733092.
- ^ Peng, Junping; Yang, Jian; Jin, Qi (2011-04-05). "An Integrated Approach for Finding Overlooked Genes in Shigella". PLOS ONE. 6 (4): e18509. Bibcode:2011PLoSO...618509P. doi:10.1371/journal.pone.0018509. ISSN 1932-6203. PMC 3071730. PMID 21483688.
- ^ Wang, Ligui; Yang, Guang; Qi, Lihua; Li, Xiang; Jia, Leili; Xie, Jing; Qiu, Shaofu; Li, Peng; Hao, RongZhang (2016-01-01). "A Novel Small RNA Regulates Tolerance and Virulence in Shigella flexneri by Responding to Acidic Environmental Changes". Frontiers in Cellular and Infection Microbiology. 6: 24. doi:10.3389/fcimb.2016.00024. ISSN 2235-2988. PMC 4782007. PMID 27014636.
- ^ Stevens J; Galyov EE; Stevens MP (2006). "Actin-dependent movement of bacterial pathogens". Nature Reviews Microbiology. 4 (2): 91–101. doi:10.1038/nrmicro1320. PMID 16415925. S2CID 30946244.
- ^ Ogawa M; Handa Y; Ashida H; Suzuki M; Sasakawa C (2008). "The versatility of Shigella effectors". Nature Reviews Microbiology. 6 (1): 11–16. doi:10.1038/nrmicro1814. PMID 18059288. S2CID 26214256.
- ^ Robbins JR; Barth AI; Marquis H; de Hostos EL; Nelson WJ; Theriot JA (1999). "Listeria monocytogenes exploits normal host cell processes to spread from cell to cell". Journal of Cell Biology. 146 (6): 1333–1350. doi:10.1083/jcb.146.6.1333. PMC 1785326. PMID 10491395.
- ^ a b c Sanada T; Kim M; Mimuro H; Suzuki M; Ogawa M; Oyama A; Ashida H; Kobayashi T; Koyama T; Nagai S; Shibata Y; Gohda J; Inoue J; Mizushima T; Sasakawa C (2012). "The Shigella flexneri effector OspI deamidates UBC13 to dampen the inflammatory response". Nature. 483 (7391): 623–6. Bibcode:2012Natur.483..623S. doi:10.1038/nature10894. PMID 22407319. S2CID 4371539.
nora https://microbenotes.com/biochemical-test-of-shigella-flexneri/
External links
- "Shigella flexneri". NCBI Taxonomy Browser. 623.
- Type strain of Shigella flexneri at BacDive - the Bacterial Diversity Metadatabase